Intrinsic Bimodality of Protein pI Distributions Our present paper overviews proteome-wide pI distributions focusing on the relationships between protein pI and subcellular localization. The localization- and function-specific differences in subcellular pI distributions have been disclosed. These studies have revealed important universal features of the whole proteome pI distributions providing insights into spatial organization of cellular proteomes. Availability of whole-genome sequences allows comparative and evolutionary studies of proteome-wide pI distributions in different organisms. The great variation of protein pI values brings about the question about the cause of this variation. Thus, protein pI is an integral property of a protein molecule fundamentally important for its characterization. Also, a database of protein pIs that were predicted using multiple available methods has been presented ( Kozlowski, 2017). 
In addition, the experimentally observed protein pI values were summarized in the experimental databases ( Hoogland et al., 2004 Bunkute et al., 2015). Some methods take into account the effect of the amino acids residues adjacent to the charged residues, such as aspartate and glutamate ( Cargile et al., 2008), effects of posttranslational modifications, such as phosphorylation and N-terminal acetylation ( Gauci et al., 2008), or effects of the presence of polyelectrolyte chains around proteins ( Srivastava et al., 2017). Various calculative algorithms have been developed for estimating protein pIs in agreement with experiments regardless of structural aspect ( Gasteiger et al., 2003 Cargile et al., 2004 Gauci et al., 2008 Maldonado et al., 2010 Audain et al., 2016). 
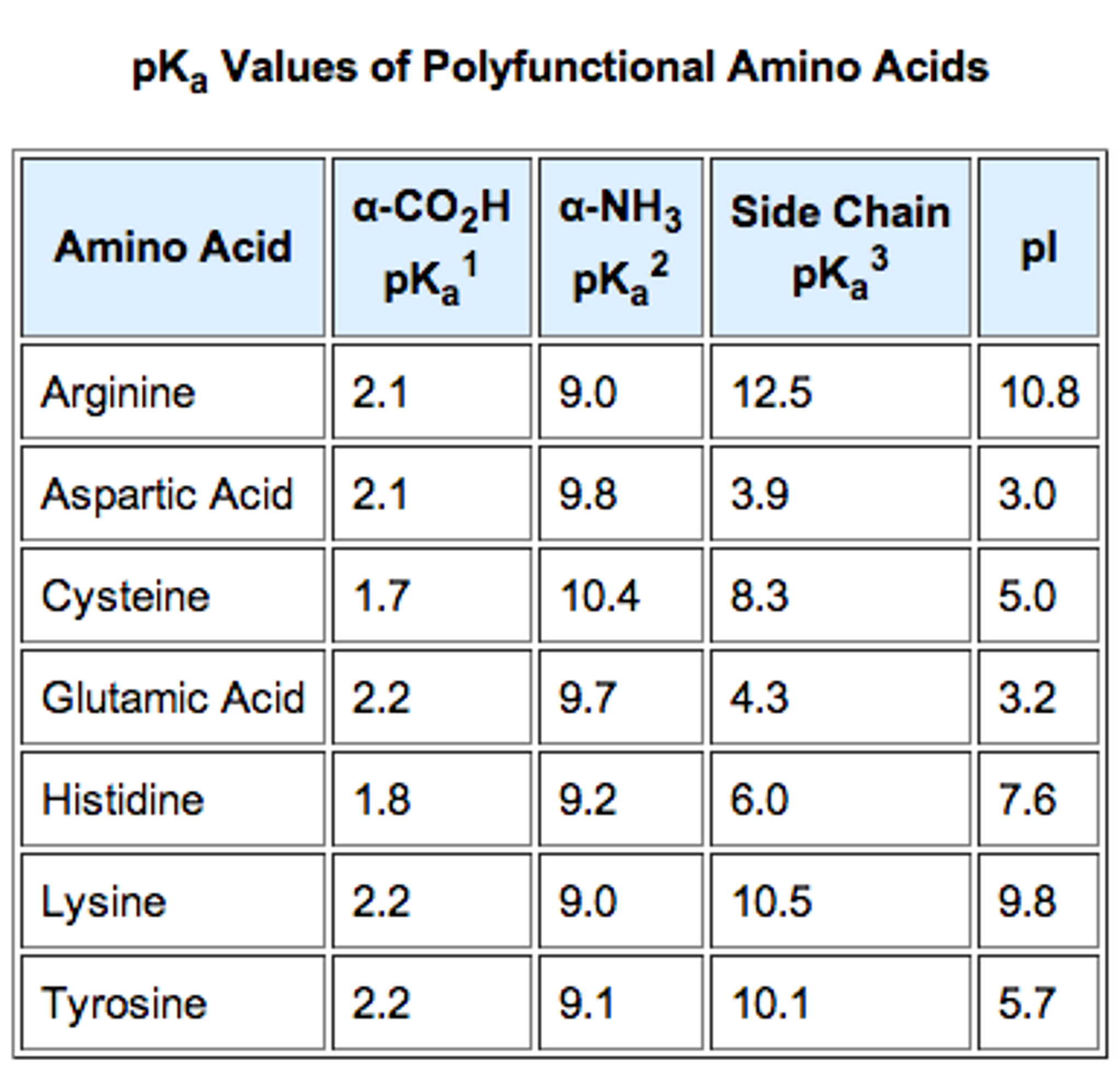
Notably, three-dimensional structure and pH of surrounding environment can influence ionizable groups and affect the net charge on the molecule significantly ( Russell and Warshel, 1985). It was demonstrated that the protein pI can be estimated based on a polypeptide sequence in close agreement with experimentally determined pI values ( Sillero and Ribeiro, 1989), and the focusing positions of proteins in immobilized pH gradients and two-dimentional gels can be reliably predicted from their amino acid composition ( Bjellqvist et al., 1993 Bjellqvist et al., 1994 Link et al., 1997). Thus, the integral property of a protein, such as protein pI, was supposed to result from discrete local acidic and basic pKas of amino acid side chains. Out of twenty common amino acids, two amino acids, aspartic acid, and glutamic acid, are negatively charged and three amino acids, lysine, arginine, and histidine, are positively charged at the neutral pH, as defined by their pKa values. Amino acid composition of a protein sequence primarily defines its pI, based on the combination of dissociation constant (pKa) values of the constituent amino acids. Hence, pI values have long been used to distinguish between proteins in methods for protein isolation, separation, purification, crystallization, etc. The protein pI varies greatly from extremely acidic to highly alkaline values ranging from about 4.0 to 12.0. Accordingly, proteins are positively charged at a pH below their pI and negatively charged at a pH above their pI. The isoelectric point (pI) of a protein is defined as the pH at which the net charge of a protein molecule is zero. We also discuss the probable factors responsible for variation of the intracellular localization-specific pI profiles. Here, we overview the multimodality of proteome-wide pI distributions in different organisms focusing on the relationships between protein pI and subcellular localization. It was further hypothesized that the observed multimodality is associated with subcellular localization-specific differences in local pI distributions. It was found that the whole-proteome distributions of protein pI values are multimodal in different species. Availability of whole-genome sequences empowers comparative studies of proteome-wide pI distributions. The protein isoelectric point (pI) can be calculated from an amino acid sequence using computational analysis in a good agreement with experimental data. 3Laboratory of Cell Signaling and Development, Faculty of Life Sciences, Kyoto Sangyo University, Kyoto, Japan.2Center for Sustainable Resource Science, RIKEN Yokohama Institute, Yokohama, Japan. 
1Department of Genetic Engineering, Faculty of Biology-Oriented Science and Technology, Kindai University, Wakayama, Japan.Tokmakov 1*, Atsushi Kurotani 2 and Ken-Ichi Sato 3 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
